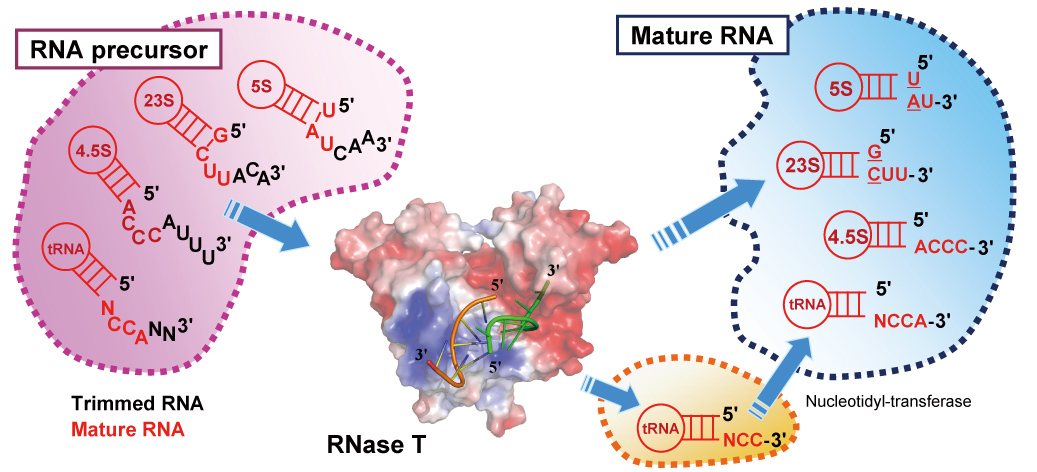
Structural basis for RNA trimming by RNase T in stable RNA 3′-end maturation
Yu-Yuan Hsiao,1, 2,Che-Chuan Yang,2, 3,Chia Liang Lin,1, 2,Jason L J Lin,2,Yulander Duh2,& Hanna S Yuan
1Institute
of Bioinformatics and Structural Biology, National Tsing Hua
University, Hsinchu, Taiwan, ROROC. 2Institute of Molecular Biology,
Academia Sinica, Taipei, Taiwan, ROROC. 3Graduate Institute of
Biochemistry and Molecular Biology, National Taiwan University, Taipei,
Taiwan, ROROC. *e-mail: hanna@sinica.edu.tw