Structural
analysis of diseaserelated TDP-43 D169G mutation: linking enhanced
stability and caspase cleavage efficiency to protein accumulation
Chien-Hao Chiang1,2, Cédric Grauffel3, Lien-Szu Wu1, Pan-Hsien Kuo1, Lyudmila G. Doudeva1,†,Carmay Lim3, Che-Kun James Shen1 & Hanna S. Yuan1
The
RNA-binding protein TDP-43 forms intracellular inclusions in
amyotrophic lateral sclerosis (ALS). While TDP-43 mutations have been
identified in ALS patients, how these mutations are linked to ALS remains
unclear. Here we examined the biophysical properties of six ALS-linked
TDP-43 mutants and found that one of the mutants, D169G, had higher
thermal stability than wild-type TDP-43 and that it was cleaved by
caspase 3 more efficiently, producing increased levels of the
C-terminal 35 kD fragments (TDP-35) in vitro and in neuroblastoma
cells. The crystal structure of the TDP-43 RRM1 domain containing
the D169G mutation in complex with DNA along with molecular dynamics
simulations reveal that the D169G mutation induces a local
conformational change in a β turn and increases the hydrophobic
interactions in the RRM1 core, thus enhancing the thermal stability of
the RRM1 domain. Our results provide the first crystal structure of
TDP-43 containing a disease-linked D169G mutation and a
disease-related mechanism showing that D169G mutant is more susceptible
to proteolytic cleavage by caspase 3 into the pathogenic C-terminal
35-kD fragments due to its increased stability in the RRM1 domain.
Modulation of TDP-43 stability and caspase cleavage efficiency could
present an avenue for prevention and treatment of TDP-43-linked
neurodegeneration.
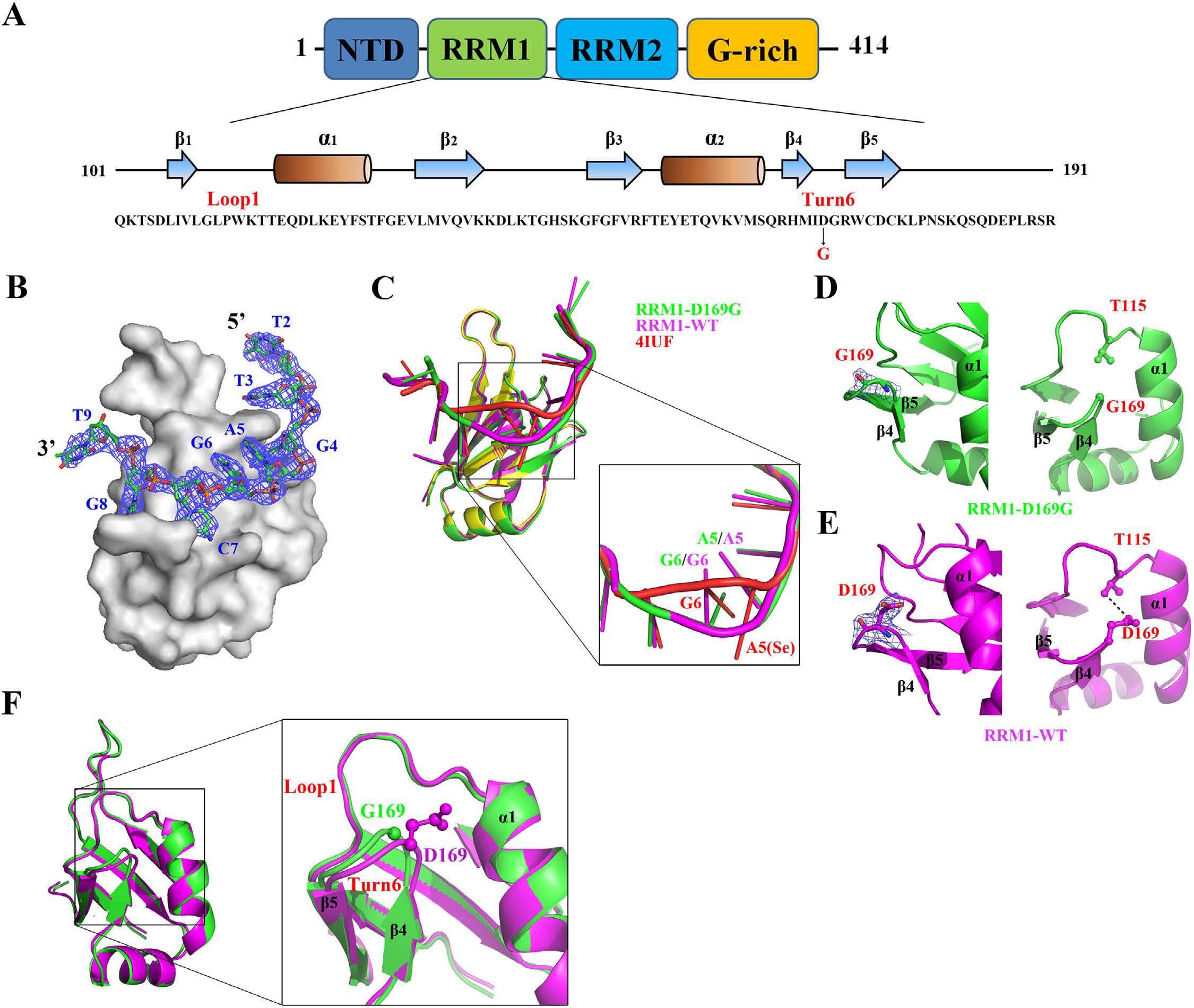
Crystal structures
of RRM1/DNA and RRM1-D169G/DNA complexes.