Research Interest
Structural
studies of nucleic acids-binding proteins in translation regulation and nucleic
acid degradation
We
are interested in a number of proteins involved
in translation regulation and DNA/RNA degradation. The overall goal is to discover
structure-based mechanisms of these proteins in nucleic acids recognition and
degradation. We use
a major tool of X-ray crystallography in combination with mutagenesis, biochemical
and biophysical approaches. Several projects of interest are listed below.
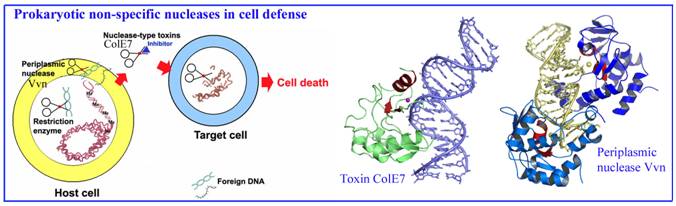
1. Bacterial nucleases in cell defense
We have been working on two types of sugar non-specific nucleases in
bacteria, including a periplasmic nuclease Vvn and a secreted toxin ColE7, both of which digest foreign nucleic acids
for cell defense. Based on our structural and biochemical analysis on Vvn
and ColE7, we have provided a solid foundation to explain how these nucleases
are inhibited and activated, how they recognize DNA without sequence
specificity and how they digest DNA to protect bacterial cells at atomic level.
Li, C., Ho, L.-I., Chang, Z.-F., Tsai, L.-C., Yang,
W.-Z. and Yuan*, H. S. (2003) DNA binding and cleavage by the periplasmic
nuclease Vvn: A novel structure with a known active site. EMBO J. 22, 4014-4025.
Hsia, K.-C., Chak, K.-F., Liang, P.-H., Cheng, Y.-S.,
Ku, W.-Y. and Yuan*, H. S. (2004) DNA binding
and degradation by the H-N-H protein ColE7. Structure
12, 205-214.
Hsia, K.-C., Li, C.-L. and Yuan*, H. S. (2005) Structural and functional
insight into the sugar-nonspecific nucleases in host defense, Curr. Opin. Struct. Biol. 15, 126-134.
Shi, Z., Chak, K.-F. and Yuan*, H. S. (2005) Identification of an essential cleavage
site in ColE7 required for import and killing cells, J. Biol. Chem. 26, 24663-24668.
Cheng, Y.-S., Shi, Z., Doudeva, L. G., Yang, W.-Z., Chak, K.-F. and Yuan*, H.
S. (2006) High-resolution crystal structure of a truncated
ColE7 translocation domain: Implications for colicin transport across
membranes. J. Mol. Biol., 356, 22-31.
Wang,
Y.-T., Yang, W.-J., Li, C.-L., Doudeva,
L. G. and Yuan*, H. S. (2007) Structural basis for sequence-dependent cleavage by nonspecific endonucleases. Nucleic Acid Res. 35, 584-594.
2. Tudor-SN in miRNA degradation and mRNA translation
regulation
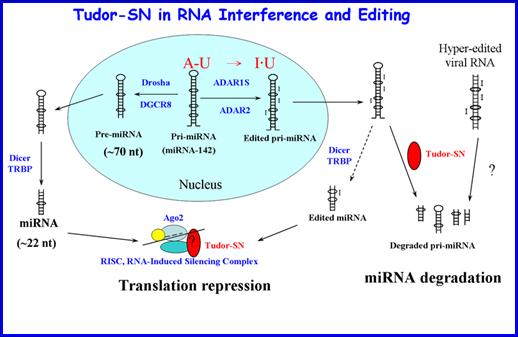
Tudor-SN
is a multifunctional protein, playing a role in transcription regulation, RNA
editing, interference and splicing. Recent studies show that Tudor-SN is a
miRNase specific for inosine-containing microRNA precursors, and it also
regulates gene expression by binding to mRNA at 3’UTR to decrease the rate of
mRNA decay. Human Tudor-SN contains five staphylococcal nuclease-like (SN) and
a tudor domains. Our structural and biochemical analysis of a truncated 64-kD
Tudor-SN shows the architecture and assembly of SN and tudor domains and also
suggests that two SN domains work together functioning as a clamp to capture
RNA substrates. Co-crystallization (with RNA), biochemical and mutagenesis
experiments are underway to further reveal the molecular basis of Tudor-SN in
mRNA recognition and miRNA cleavage.
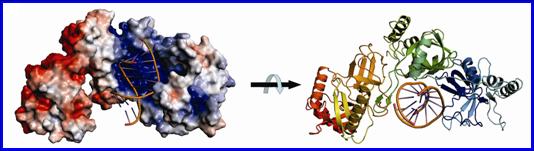
Structural model of a
64-kD Tudor-SN bound to double-stranded RNA.
Our biochemical and structural data suggest that tandem repeats of SN domains in
Tudor-SN work together to capture RNA substrates.
References:
Li,
C.-L., Yang, W.-Z., Chen, Y.-P. and Yuan*, H. S. (2008) Structural and
functional insights into human Tudor-SN, a key component linking RNA
interference and editing. Nucleic Acid Res. 36, 3579-3589.
3. PNPase in mRNA degradation
PNPase
(polynucleotide
phosphorylase) is an important enzyme responsible for mRNA turnover from
3’- to 5’-end in bacteria. It shares a similar structural organization to
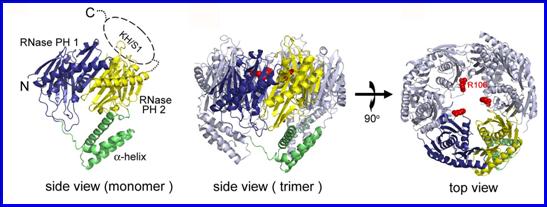
eukaryotic exosome core complexes. Our structural and biochemical study on E. coli PNPase shows that the trimeric structure of a KH/S1-truncated
PNPase is more expanded, containing a slightly wider central RNA-binding
channel than that of the wild-type PNPase. This result suggests that the KH/S1
domain is involved not only in RNA binding but it also helps PNPase to assemble
into a more compact trimer. This finding is
likely a general phenomenon since only bacterial PNPase and archaeal exosomes
with constricted channels are efficient enzymes in RNA degradation. We also
study the structures and biochemical properties of human exosome component
proteins and mitochondrial PNPase to further characterize the
structure-and-function relationship of PNPase in mRNA binding and degradation.
Reference:
Shi, Z., Yang, W.-Z., Lin-Chao, S., Chak, K.-F. and Yuan*, H. S. Crystal structure of Escherichia coli PNPase: central channel residues are involved in processive
RNA degradation. RNA (in press).
4. Apoptotic nucleases in DNA degradation.
Apoptotic
nucleases are activated for chromosomal
DNA fragmentation during
apoptosis. Inactivation of these apoptotic nucleases produces undigested
DNA and is related to a number of autoimmune disorders. We analyze the
biochemical properties and crystal structures of a number of apoptotic
nucleases to address the function of these nucleases in normal versus apoptotic
cells. Recently, we determined the crystal structure of a C. elegans cell-death-related nuclease 4 (CRN-4). The biochemical,
structural, and functional assays consistently suggest that the C-terminal
novel-fold Zn-domain of CRN-4 is involved in DNA binding and the N-terminal
nuclease domain is responsible for DNA degradation. This study therefore
provides new insights into the DEDDh family of nucleases in chromosomal DNA
fragmentation in apoptosis.
We also analyze
the biochemical and structural features of several apoptotic proteins and
nucleases that interact with CRN-4 to form a degradeosome in apoptosis,
including CPS-6 (human Endo G homologue), WAH-1 (AIF), CRN-5 (Rrp46) and
Cyp-13. The long-term goal of this research is to decipher the working
mechanism of the degradeosome in DNA fragmentation during apoptosis.
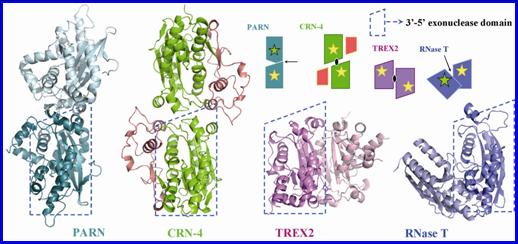
Crystal structure of CRN-4 and Comparison of the different domain arrangement in dimeic DEDDh family proteins.
CRN-4 dimerizes in a different mode as compared to PARN, TREX2 and RNase T.
Reference:


